Data Networks using BioCASe
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
VH/de |  |
A network of primary biodiversity repositories
The Biological Collection Access Service (BioCASe) is a transnational network of primary biodiversity repositories. It links together specimen data from natural history collections, botanical/zoological gardens and research institutions worldwide with information from huge observation databases. The aim is to make the world's data on biodiversity data freely and universally accessible on the Internet through data portals and web services, a goal that BioCASe shares with related initiatives such as the Global Biodiversity Information Facility (GBIF) and Integrated Digitized Biocollections (iDigBio). In the past years, BioCASe has developed into a widely accepted standard for data sharing and has laid the foundations for several thematically - both taxonomically and geographically - specialized networks (see box to the right). Read more
 The BioCASe secretariat is based at the Botanic Garden and Botanical Museum Berlin and will be happy to answer any questions regarding BioCASe.
The BioCASe secretariat is based at the Botanic Garden and Botanical Museum Berlin and will be happy to answer any questions regarding BioCASe.

BioCASe is the GBIF Participant Node for the Consortium of European Taxonomic Facilities.

About 23 million dried plants, fungi and algae are safely stored in more than 70 German herbaria. One million of them can be explored now in the German Virtual Herbarium. More than 300,000 of the records come with at least one digital image. The specimens can be viewed as a list, map or in a gallery and can be filtered by a multitude of search criteria (more on the features can be found in this news item).
The Virtual Herbarium Germany is curated by the Botanic Garden Berlin of Freie Universität Berlin and is based on the Hosted Portal technology provided by the Global Biodiversity Information Facility. All the 22 collections accessible through the Virtual Herbarium use the BioCASe Provider Software to publish their specimens to GBIF and other biodiversity networks.

A new version of BioCASe has been published that runs on Python 3. For this, the software underwent a thorough overhaul, which included a lot of code cleaning and bug fixing.
The Python 3 version is up to 50% faster due to code removal/re-implementation and Python optimizations (tested on MySQL and SQL Server databases). External dependencies have been reduced, which makes installation way easier. Last but not least, BioCASe now deals better with unexpected control characters in the sources databases, which caused a lot of pain in the previous versions.

The CETAF Earth Science Group (ESG) has just announced that the first Minimum Viable Product of GeoCASe 2.0 is online and ready for use. GeoCASe is using the BioCASe Provider Software for publishing data from the institutional source databases to the data portal.
GeoCASe stands for Geoscience Collections Access Service, which was created in 2007. Now the completely revamped interface allows users to more efficiently find data on minerals, rocks, meteorites and fossils held in multiple European natural history museums and research institutions. Quick search capabilities, data download options, images and map views, and detailed specimen pages are among the new key functions of the portal.

The latest version of the BioCASe Provider Software includes a template for the botanical garden and herbarium management system IrisBG. With this, you can easily connect your database to BioCASe if you're using the cloud-based version of IrisBG - all you need to do is ask the IrisBG team/Compositae to grant access to your database and enter the credentials you got from them into BioCASe.
Another new feature was added by Peter Grobe to BioCASes Local QueryTool: The details page will now show a preview for certain multimedia types, including images, videos and sound files.The release contains several fixes and other small changes; as always, the complete list can be found in the version history.
A version of the BioCASe Provider Software has been released. Apart from numerous smaller changes, it can now be installed as a Docker container. Docker is virtualization software that allows easy deployment and maintenance of software packages. Instead of installing a web server, Python packages and BioCASe consecutively, the whole suite can be easily installed from a Docker images with a single command. To learn about how to install BioCASe from a Docker image, see the installation guide.
Another new feature are filtered exports: They allow a subset of the records published in a data source to be stored in an XML archive. The subset can be defined as a BioCASe filter - that is, by using operators such as equals, like, lessThan/greatherThan and isNotNull ony any ABCD concept mapped.
For a complete list of changes and fixes, see the version history.
Two more regional biodiversity networks based on BioCASe have been created recently:
The Biodiversity Network of Mozambique (BioNoMo) currently consists of four data providers, with more to be added in the near future. The data portal is still a pilot installation, but already equipped with a central cache database storing the data provided in the ABCD 2.06 data standard.
The Biodiversity National Network of Albania (BioNNA) is providing an important entry point for any initiative aiming to support decisions related to biodiversity conservation and ecosystem management with a data portal based on distributed queries.

One of the main uses cases for the BioCASe Provider Software is data provision to the Global Biodiversity Information Facility (GBIF). Currently roughly 35 million specimen and observation records are fed into the GBIF network, which can be accessed and downloaded through the GBIF data portal.
The newly created GBIF data usage page displays usage statistics for those BioCASe-mediated information. It lists the number of records downloaded through GBIF, grouped by provider, year and month, as well as the total number of BioCASe records contained in these downloads. The numbers are updated weekly.

The new version of the BioCASe Provider Software is now able to publish data in the LIDO format (Lightweight Information Describing Objects). This standard is widely used in the museum community. With 1.261 data elements and attributes (3.041 including GML fields) being even heavier than ABCD, LIDO is able to support the full range of descriptive information about museum objects. It can be used for all kinds of objects, e.g. art, architecture, cultural history, history of technology, and natural history. With the BioCASe Provider Software now supporting LIDO, data from any SQL-based database can be easily exported into LIDO.
The mapping editor has undergone several changes to better reflect the characteristics of the schema used, for example mandatory elements or nodes. XML archiving has been improved to use less memory now and tolerate temporary downtimes of the database to be archived. Other improvements affect DarwinCore archives and the modules for connecting to Microsoft SQL Server. For a complete list of changes and fixes, please refer to the version history.

The German Virtual Herbarium, abbreviated VH/de, is a new data portal facilitating access to the plant collections records of German herbaria. These collections house more than 15 million specimens of vascular plants, mosses and macro algae and have an extraordinary value as archives and documentation of species occurrences over time and space. They are essential for research on systematics, taxonomy, abundance of plants and on changes of the flora of our planet. The data portal is based on the BioCASe Portal Software.
The new data portal offers a common entrance point to the digitally available parts of these collections. Currently, more than 530,000 herbarium specimens are accessible at http://vh.gbif.de. Up to now, fourteen herbaria from Germany have joined, with more to follow. VH/de is being launched with collections from Berlin (B), Bremerhaven (BRM), Frankfurt/Main (FR), Görlitz (GLM), Göttingen (GOET), Halle (HAL), Hamburg (HBG), Jena (JE), Konstanz (KONL), Mainz (MJG, MNHM), München (M, MSB) and Regensburg (REG). All of the partners use the BioCASe Provider Software for feeding their data into the Virtual Herbarium.
The common access point for several herbaria allows comparative research on specimens from different institutions. Upon request, the German Virtual Herbarium always shows the latest data, since it retrieves records directly from the data provider before display. High-resolution images allow close-up scrutiny of the virtual specimens.

With the latest version of the BioCASe Provider Software, it is now possible to publish data from Excel spreadsheets. Just like with any other database system, you can map a data standard like ABCD to different spreadsheets and columns. For small collections, this provides an easy way of publishing data to biodiversity networks without the need of database imports.
Moreover, the QueryTool of the Provider Software can now use the libxml/libxslt packages instead of 4SuiteXML for stylesheet transformation. This allows BioCASe to be used with Python versions 2.6 and 2.7, for which the 4SuiteXML package is not available. Several smaller changes and fixes are part of this release; for a complete list, please refer to the version history.

With the latest version of the BioCASe Provider Software, it becomes easier to update new versions. The setup script is now able to import the configuration of an existing BioCASe installation, including all data sources configured. An update hint on the start page will inform administrators of the installation of available updates.
Additional data items have been added to the DarwinCore archives produced by BioCASe, reducing the amount of information lost in the transformation from ABCD to DarwinCore. Firebird and Sybase databases are supported now, paging has been optimized for Oracle. Several smaller changes and fixes are part of this release; for a complete list, please refer to the version history.
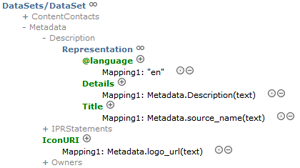
In the latest version of the BioCASe Provider Software, the mapping editor got a facelift. The concept tree, which can be quite large for complex data schemas as ABCD, is now collapsible. This allows nodes that are of no interest to be collapsed during the mapping process for a better overview of the tree.
The XML archiving now shows the progress indicator in percent, which will allow the user to better estimate the time required for completing the process. The new OFFSET clause introduced in SQL Server 2012 is now supported for better performance when huge datasets are paged. For a complete list of changes, see the version history.
In the first volume of the Biodiversity Data Journal (BDJ) a new community peer-reviewed, open-access publishing platform for biodiversity-related data , an article on the BioCASe Monitor Service (BMS) has been published. This new tool for monitoring installations of the BioCASe Provider Software has been developed by a team at Natural History Museum Berlin (MfN) and Royal Museum of Central Africa Tervuren (RMCA).
It is released under the Creative Commons CC0 license and can be used to monitor networks of BioCASe data providers. It will aid the network manager in monitoring the availability of the providers, in checking the data mappings for completeness and plausibility and in keeping track of the providers progress in data provision (number of records published). Currently, it is used by two projects, OpenUp! and GBIF-D, the German GBIF node. More information on the tool as well as links to documentation and download location can be found in the BDJ article.
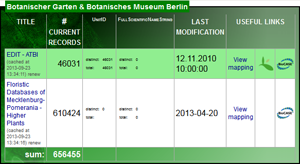
BioCASe allows publishing several datasets with a single web service, all datasets sharing the same technical access point. With the latest version of the BioCASe Provider Software, administrators can restrict XML archiving to single datasets within a web service, allowing the selective creation or update of XML and DarwinCore Archives. This improves performance significantly, paving the way for frequent (automated) updates of archives for continuously or regularly updated datasets.
A new optional attribute "expires has been added to the Dataset Inventory. Based on a validity period set by the web service administrator, archives can be flagged automatically as outdated by the Provider Software. The release contains several fixes and other small changes; the complete list can be found in the version history.
In the latest version 3.4 of the BioCASe Provider Software, the XML archiving features have been re-implemented completely and aligned with GBIFs new dataset-aware Registry. Now each dataset published by a BioCASe web service will end up in a separate XML archive, storing all information published by the web service. GBIF will use the newly introduced BioCASe dataset inventory to discover these archives and use them for faster indexing. If desired, individual archives or the complete list of XML archives can be transformed into DarwinCore archives.
In addition, the archiving process has been streamlined and should be significantly faster on most installations; instead of using the CGI interface, the BioCASe libraries are now called directly. Dispensable formatting has been removed, reducing the size of XML documents stored in the archives by about one third. The archiving interface underwent a a comprehensive redesign.
The new release contains several fixes and other small changes; the complete list can be found in the version history.
A version of the BioCASe Provider Software with smaller bug fixes and changes has been released. Now it is possible to transfer the mappings from one schema to another schema with the namespace. This allows, for example, upgrading from ABCD 2.06 to one of the extensions ABCD-EFG or ABCD-DNA or vice versa. More upgrade paths have been added.
For a complete list of changes and fixes, see the version history.
A new version of the BioCASe Provider Software has been released. It now supports the archiving processes to be triggered through a deep link, without using the configuration tool interface. This allows archiving to be scheduled; for example, a DarwinCore Archive could be created monthly for being harvested by GBIF or any other biodiversity network. Both XML and DarwinCore Archives can be created through the deep link.
Several other changes have been implemented; for example, the ordering of Scan request results has been changed in order to increase compatibility with widely used indexing approaches. For a complete list of changes, see the version history.
Version 3.2 of the BioCASe Provider Software has been released. It allows easy creating of DarwinCore Archives from an ABCD web service with just one click in the user interface. For special interest networks relying on the full richness of ABCD concepts, ABCD archives or other XML archives can be created in a similar fashion. Both archive types can be used to speed up and ease harvesting and indexing processes.
For a complete list of changes, see the version history.
A new version of the BioCASe Provider Software has been released. The focus for improvements has been on the new XML archiving feature: The web interface has been overhauled and reacts faster now. If it is kept open during the archiving process, less load is put on the server for showing the progress log. Several smaller bugs have been fixed.
For a complete list of changes, see the version history.
Shortly before Easter, a new version of the BioCASe Provider Software has been released. It comprises mainly bug fixes. Compatibility with Internet Information Server has been improved; if SQL Server is used, the configuration tool now provides drop-down lists for table and column names.
A complete list of bug fixes can be found in the version history.
A new major version of the BioCASe Provider Software has been released. As an important new feature, version 3 can now create XML archives that store all information published by a BioCASe web service in a single file. If required, these archives can be transformed into DarwinCore Archives in a subsequent step.
In addition, a number of smaller changes and optimizations have been implemented. Metadata retrieval is now also supported for Oracle databases, which allows an easier configuration of the according web services. For a complete list of changes, see the version history.
A new version of the Provider Software has been released. Now most configuration pages feature a Help link that points directly to the respective tutorial of the new PyWrapper documentation wiki. Moreover, it contains a number of smaller bug fixes, please see the version history for a complete list.

The new PyWrapper documentation wiki is now online at http://wiki.bgbm.org/bps. It has been rewritten completely from scratch for the latest version of the BioCASe Provider Software (2.6.0).
It features several step-by-step tutorials on installation, configuration, debugging, on preparing a database for publication with BioCASe, a beginners guide, a short documentation of ABCD, and a list of frequently asked questions. Many pages in the upcoming version of the Provider Software will include a Help link that points directly to the respective page in the online documentation.
Version 2.6 of the Provider Software has been released. It features major changes in the user interface for the configuration tool that should ease the setup of the BPS considerably. Table setup has been completely overhauled; table and column names will now be retrieved from the database and shown in drop-down-boxes for commonly used DBMS. The mapping editor also uses drop-down-boxes now.
If MySQL or Postgres are used, the Provider Software now takes advantage of the OFFSET SQL clauses for serving Search requests, which increases performance for harvesting applications. Last but not least, a number of bugs have been fixed. See the Version History for a complete list of changes.
The OpenUP! project launched in March will use BioCASe Technology to connect millions of multimedia objects stored in natural history collections to the European network Europeana, thereby making them accessible to the public in a high quality.
In order to build the required expertise in the associated project partners, a two day hands-on workshop was held on May 30th and 31st at the Botanic Museum in Berlin to teach the installation and configuration of the BioCASe Provider Software. 17 Participants from 9 countries will now use their knowledge to set up BioCASe installations in their own institutions.
For older news items please see the news archive.
